dwctaxon is all about handling taxonomic data in the Darwin Core (DwC) taxon format.
But what is DwC?
DwC is a standard for biodiversity data
According to the official documentation,
Darwin Core is a standard maintained by the Darwin Core Maintenance Interest Group. It includes a glossary of terms (in other contexts these might be called properties, elements, fields, columns, attributes, or concepts) intended to facilitate the sharing of information about biological diversity by providing identifiers, labels, and definitions. Darwin Core is primarily based on taxa, their occurrence in nature as documented by observations, specimens, samples, and related information.
(emphasis added)
The “terms” referred to are typically encountered as columns in
rectangular data (spreadsheets), such as scientificName
(scientific name of a taxon), lifeStage (life stage of an
organism when it was observed), etc. By providing a controlled
vocabulary and clear definitions of terms, DwC greatly facilitates
collection and sharing of biological data. For example, the Global Biodiversity Information Facility
(GBIF), which synthesizes biodiversity data on a global scale, uses DwC.
In practice, a given set of DwC data are contained in an archive (zip file) including multiple spreadsheets (CSV files) and XML files with additional metadata. The spreadsheets typically include datasets like occurrences, taxonomy, and collection events.
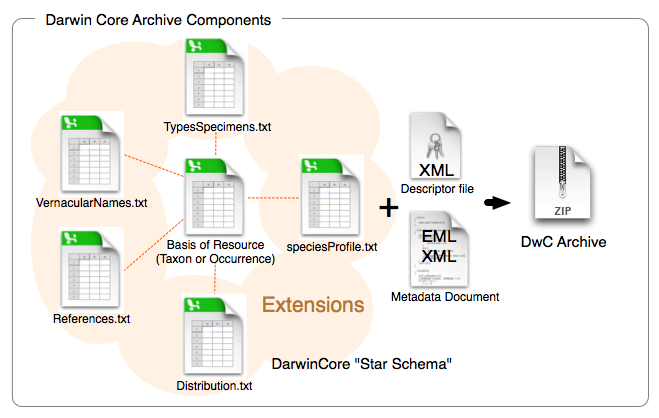
While other parts of DwC such as organism and occurrence data are clearly important, they are out of the scope of dwctaxon, which only focuses on taxonomic data.
Features of the DwC taxon format
Most of the terms used in the DwC format for taxonomic data (“DwC taxon”) should be familiar to biologists. Here is a simple example mapping taxonomic data on the left to their DwC terms on the right for the genus Sarda:
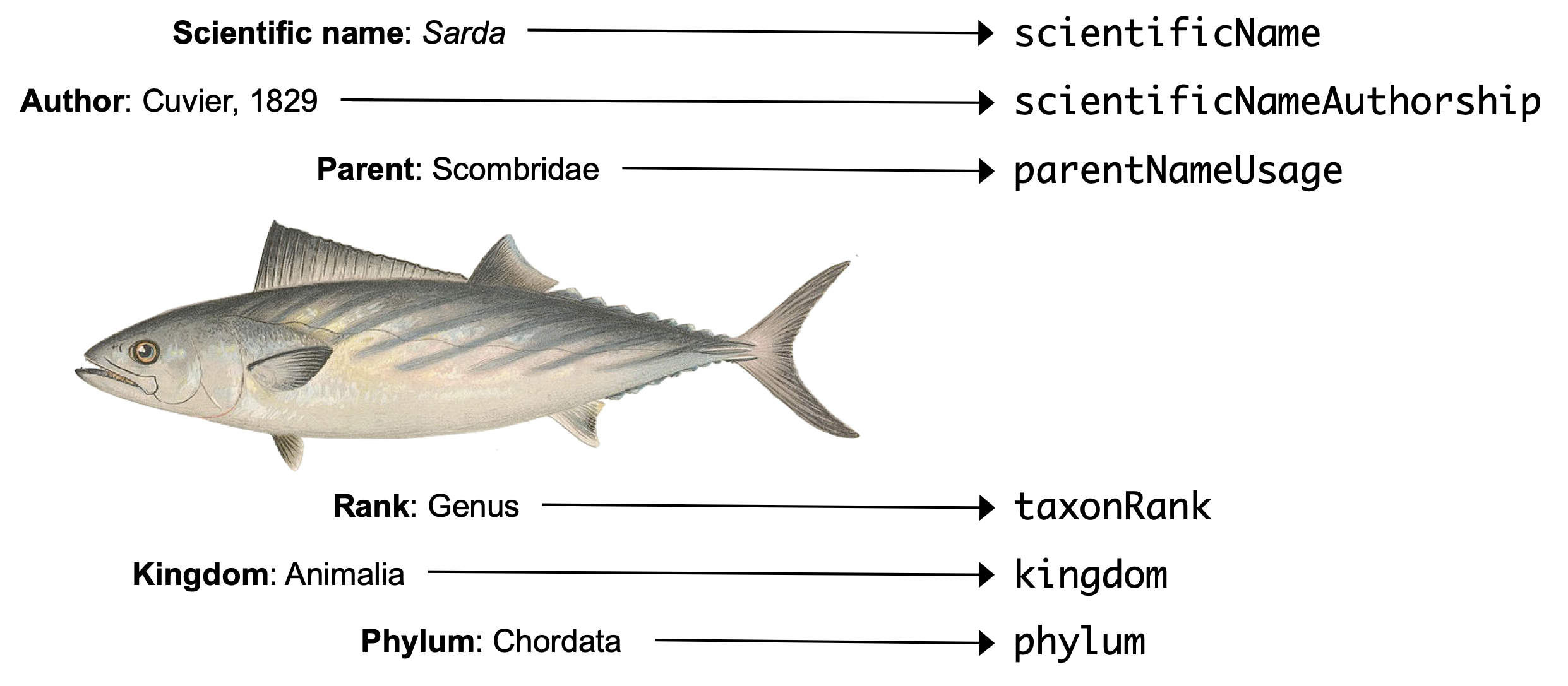

However, there are some peculiarities that are good to be familiar with as follows.
Vertically oriented
The Linnaean system of taxonomy organizes taxa into a hierarchy, so we may be used to working with taxonomic data in “wide” format where each row is a species, like this:
| species | genus | family | order |
|---|---|---|---|
| Crepidomanes minutum | Crepidomanes | Hymenophyllaceae | Hymenophyllales |
Indeed, in DwC taxon, taxonomic levels above species like
genus, family, and order are
valid terms and may be used.
However, species is not a valid DwC
term. That is because each row of a DwC taxonomic database is a single
scientific name of any rank, not just species. So it is
typical for data to be oriented vertically (“long” format):
| taxonRank | scientificName |
|---|---|
| species | Crepidomanes minutum |
| genus | Crepidomanes |
| family | Hymenophyllaceae |
| order | Hymenophyllales |
And since genus, family, etc. are valid DwC
terms, these can also be included (when applicable):
| taxonRank | scientificName | genus | family | order |
|---|---|---|---|---|
| species | Crepidomanes minutum | Crepidomanes | Hymenophyllaceae | Hymenophyllales |
| genus | Crepidomanes | Crepidomanes | Hymenophyllaceae | Hymenophyllales |
| family | Hymenophyllaceae | NA | Hymenophyllaceae | Hymenophyllales |
| order | Hymenophyllales | NA | NA | Hymenophyllales |
Machine and human friendly
If you browse through the DwC taxon terms, you will notice many
pairs of similar terms such as acceptedNameUsage and
acceptedNameUsageID, parentNameUsage and
parentNameUsageID, etc. These each are used for similar
purposes, but one is a value that is easy for humans to understand while
the other is useful for machines (computer programs).
For example, acceptedNameUsage is the accepted name of a
synonym (e.g., Picea abies (L.) H. Karst as the accepted name
of Pinus abies L.), and acceptedNameUsageID is the
unique ID (typically, taxonID) of the accepted name
(typically some short sequence of letters and numbers, but this depends
on the dataset).
This makes the data format somewhat redundant, but it is also easier
for a human to parse if they can see the actual accepted name of a
synonym immediately, instead of having to look it up by
taxonID. On the other hand, scientificName can
include duplicates (in rare cases if the same name was published twice,
etc.), so referring to an accepted name by its unique ID is safer and no
problem for a computer.
We can see how this works in the example dataset that comes with
dwctaxon, dct_filmies:
library(dwctaxon)
head(dct_filmies)
#> # A tibble: 6 × 5
#> taxonID acceptedNameUsageID taxonomicStatus taxonRank scientificName
#> <chr> <chr> <chr> <chr> <chr>
#> 1 54115096 NA accepted species Cephalomanes atrovirens Presl
#> 2 54133783 54115097 synonym species Trichomanes crassum Copel.
#> 3 54115097 NA accepted species Cephalomanes crassum (Copel.) M. G. Price
#> 4 54133784 54115098 synonym species Trichomanes densinervium Copel.
#> 5 54115098 NA accepted species Cephalomanes densinervium (Copel.) Copel.
#> 6 54133786 54115100 synonym species Cephalomanes curvatum (J. Sm.) V. D. BoschHere, Trichomanes crassum Copel. is a synonym of
Cephalomanes crassum (Copel.) M. G. Price (notice how the
acceptedNameUsageID of Trichomanes crassum Copel.
matches the taxonID of Cephalomanes crassum
(Copel.) M. G. Price).
In this dataset, only acceptedNameUsageID is used, but
it would be valid to add a column with acceptedNameUsage.
To learn more about how to do so, please see the
vignette("editing").
Extensible
There are many terms listed in the DwC taxon documentation – 37 by my
count! However, it is unlikely a given taxonomic database uses all of
them; in fact, most that I’ve encountered only use a subset of the
terms, and there are none that are strictly required. But in practice
you typically want at least scientificName (scientific name
of the taxon, including author if known) and taxonID (a
unique identifier for each row in the dataset).
Furthermore, some of the terms are likely to have restricted
vocabularies. For example, a given dataset may only use a limited set of
words to describe taxonomicStatus like “accepted”,
“synonym”, and “doubtful”. This is in contrast to a term that could be
(nearly) anything, like scientificName. DwC does not
provide any official set of vocabularies; it is left to the database
manager to determine that. One feature of dwctaxon is to verify that
only the values you want to allow are used for a given term. To learn
more about that, please see vignette("validation").
These qualities make the DwC taxon format flexible, so it can meet the needs of the dataset at hand. The dwctaxon functions try to provide sensible defaults, but they may need to be adjusted appropriately.
