Following the template in OpenAlex’s oa-percentage tutorial, this vignette uses openalexR to answer:
How many of recent journal articles from the University of Pennsylvania are open access? And how many aren’t?
We first need to find the openalex.id
for University of Pennsylvania. We can do this by fetching for the
institutions entity and put “University of
Pennsylvania” in display_name or
display_name.search:
oa_fetch(
entity = "inst", # same as "institutions"
display_name.search = "\"University of Pennsylvania\""
) %>%
select(display_name, ror) %>%
knitr::kable()| display_name | ror |
|---|---|
| University of Pennsylvania | https://ror.org/00b30xv10 |
| California University of Pennsylvania | https://ror.org/01spssf70 |
| Hospital of the University of Pennsylvania | https://ror.org/02917wp91 |
| University of Pennsylvania Health System | https://ror.org/04h81rw26 |
| Indiana University of Pennsylvania | https://ror.org/0511cmw96 |
| University of Pennsylvania Press | https://ror.org/03xwa9562 |
| Cheyney University of Pennsylvania | https://ror.org/02nckwn80 |
We will use the first ror, 00b30xv10, as one of the filters for our query.
Alternatively, we could go to the autocomplete endpoint at https://explore.openalex.org/ to search for “University of Pennsylvania” and find the ror there!
All other filters are straightforward and explained in detailed in
the original jupyter notebook tutorial.
The only difference here is that, instead of grouping by
is_oa, we’re interested in the “trend” over the years, so
we’re going to group by publication_year, and perform the
query twice, one for is_oa = "true" and one for
is_oa = "false" .
open_access <- oa_fetch(
entity = "works",
institutions.ror = "00b30xv10",
type = "article",
from_publication_date = "2012-08-24",
is_paratext = "false",
is_oa = "true",
group_by = "publication_year"
)
closed_access <- oa_fetch(
entity = "works",
institutions.ror = "00b30xv10",
type = "article",
from_publication_date = "2012-08-24",
is_paratext = "false",
is_oa = "false",
group_by = "publication_year"
)
uf_df <- closed_access %>%
select(- key_display_name) %>%
full_join(open_access, by = "key", suffix = c("_ca", "_oa"))
uf_df
#> key count_ca key_display_name count_oa
#> 1 2018 4451 2018 5618
#> 2 2015 4293 2015 4690
#> 3 2014 4213 2014 4531
#> 4 2025 4202 2025 8043
#> 5 2019 4162 2019 6453
#> 6 2022 4154 2022 7614
#> 7 2020 4151 2020 8020
#> 8 2013 4144 2013 4435
#> 9 2021 4073 2021 8147
#> 10 2016 3998 2016 4909
#> 11 2024 3914 2024 8269
#> 12 2017 3913 2017 5180
#> 13 2023 3468 2023 8448
#> 14 2026 1572 2026 2526
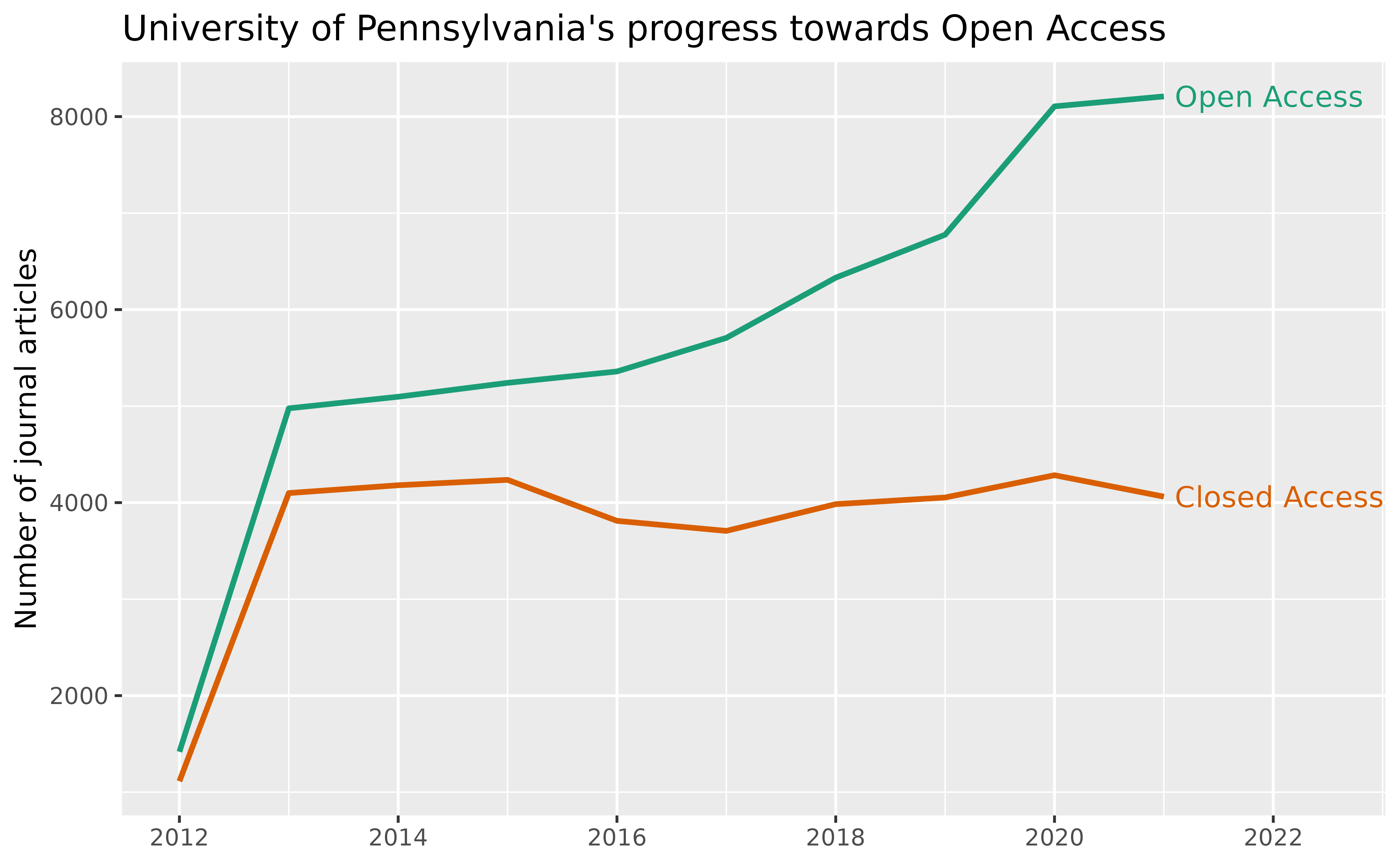
#> 15 2012 1291 2012 1091Finally, we compare the number of open vs. closed access articles over the years:
uf_df %>%
filter(key <= 2021) %>% # we do not yet have complete data for 2022 and after
pivot_longer(cols = starts_with("count")) %>%
mutate(
year = as.integer(key),
is_oa = recode(
name,
"count_ca" = "Closed Access",
"count_oa" = "Open Access"
),
label = if_else(key < 2021, NA_character_, is_oa)
) %>%
select(year, value, is_oa, label) %>%
ggplot(aes(x = year, y = value, group = is_oa, color = is_oa)) +
geom_line(size = 1) +
labs(
title = "University of Pennsylvania's progress towards Open Access",
x = NULL, y = "Number of journal articles") +
scale_color_brewer(palette = "Dark2", direction = -1) +
scale_x_continuous(breaks = seq(2010, 2024, 2)) +
geom_text(aes(label = label), nudge_x = 0.1, hjust = 0) +
coord_cartesian(xlim = c(NA, 2022.5)) +
guides(color = "none")