All the tests were done on an Arch Linux x86_64 machine with an Intel(R) Core(TM) i7 CPU (1.90GHz).
Empirical likelihood computation
We show the performance of computing empirical likelihood with
el_mean(). We test the computation speed with simulated
data sets in two different settings: 1) the number of observations
increases with the number of parameters fixed, and 2) the number of
parameters increases with the number of observations fixed.
Increasing the number of observations
We fix the number of parameters at
,
and simulate the parameter value and
matrices using rnorm(). In order to ensure convergence with
a large
,
we set a large threshold value using el_control().
library(ggplot2)
library(microbenchmark)
set.seed(3175775)
p <- 10
par <- rnorm(p, sd = 0.1)
ctrl <- el_control(th = 1e+10)
result <- microbenchmark(
n1e2 = el_mean(matrix(rnorm(100 * p), ncol = p), par = par, control = ctrl),
n1e3 = el_mean(matrix(rnorm(1000 * p), ncol = p), par = par, control = ctrl),
n1e4 = el_mean(matrix(rnorm(10000 * p), ncol = p), par = par, control = ctrl),
n1e5 = el_mean(matrix(rnorm(100000 * p), ncol = p), par = par, control = ctrl)
)Below are the results:
result
#> Unit: microseconds
#> expr min lq mean median uq max neval
#> n1e2 435.469 476.8905 522.720 500.290 557.626 757.258 100
#> n1e3 1267.758 1446.8045 1580.421 1547.079 1673.818 2709.245 100
#> n1e4 11393.587 13138.8710 15475.314 15941.224 16947.373 22562.019 100
#> n1e5 184991.234 221522.1545 255491.712 251015.169 275534.833 410394.558 100
#> cld
#> a
#> a
#> b
#> c
autoplot(result)
#> Warning: `aes_string()` was deprecated in ggplot2 3.0.0.
#> ℹ Please use tidy evaluation idioms with `aes()`.
#> ℹ See also `vignette("ggplot2-in-packages")` for more information.
#> ℹ The deprecated feature was likely used in the microbenchmark package.
#> Please report the issue at
#> <https://github.com/joshuaulrich/microbenchmark/issues/>.
#> This warning is displayed once per session.
#> Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
#> generated.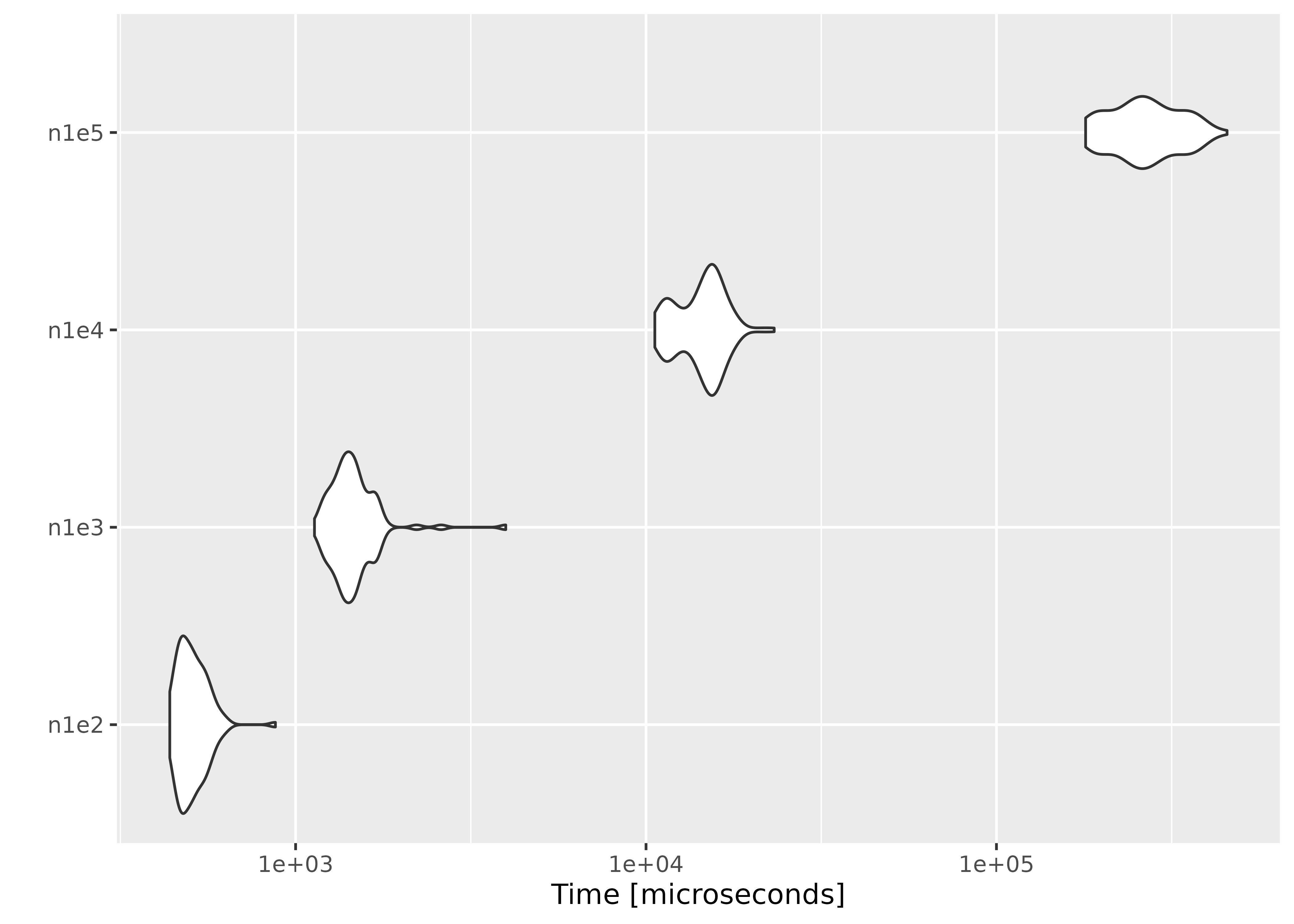
Increasing the number of parameters
This time we fix the number of observations at , and evaluate empirical likelihood at zero vectors of different sizes.
n <- 1000
result2 <- microbenchmark(
p5 = el_mean(matrix(rnorm(n * 5), ncol = 5),
par = rep(0, 5),
control = ctrl
),
p25 = el_mean(matrix(rnorm(n * 25), ncol = 25),
par = rep(0, 25),
control = ctrl
),
p100 = el_mean(matrix(rnorm(n * 100), ncol = 100),
par = rep(0, 100),
control = ctrl
),
p400 = el_mean(matrix(rnorm(n * 400), ncol = 400),
par = rep(0, 400),
control = ctrl
)
)
result2
#> Unit: microseconds
#> expr min lq mean median uq max neval
#> p5 711.149 774.203 867.1492 816.441 876.220 4686.820 100
#> p25 2902.883 2983.633 3084.4227 3028.565 3095.715 6833.586 100
#> p100 23114.201 25948.242 28258.3973 26421.422 30745.145 48674.835 100
#> p400 257745.096 283658.526 321392.8748 306667.650 350381.443 479396.343 100
#> cld
#> a
#> a
#> b
#> c
autoplot(result2)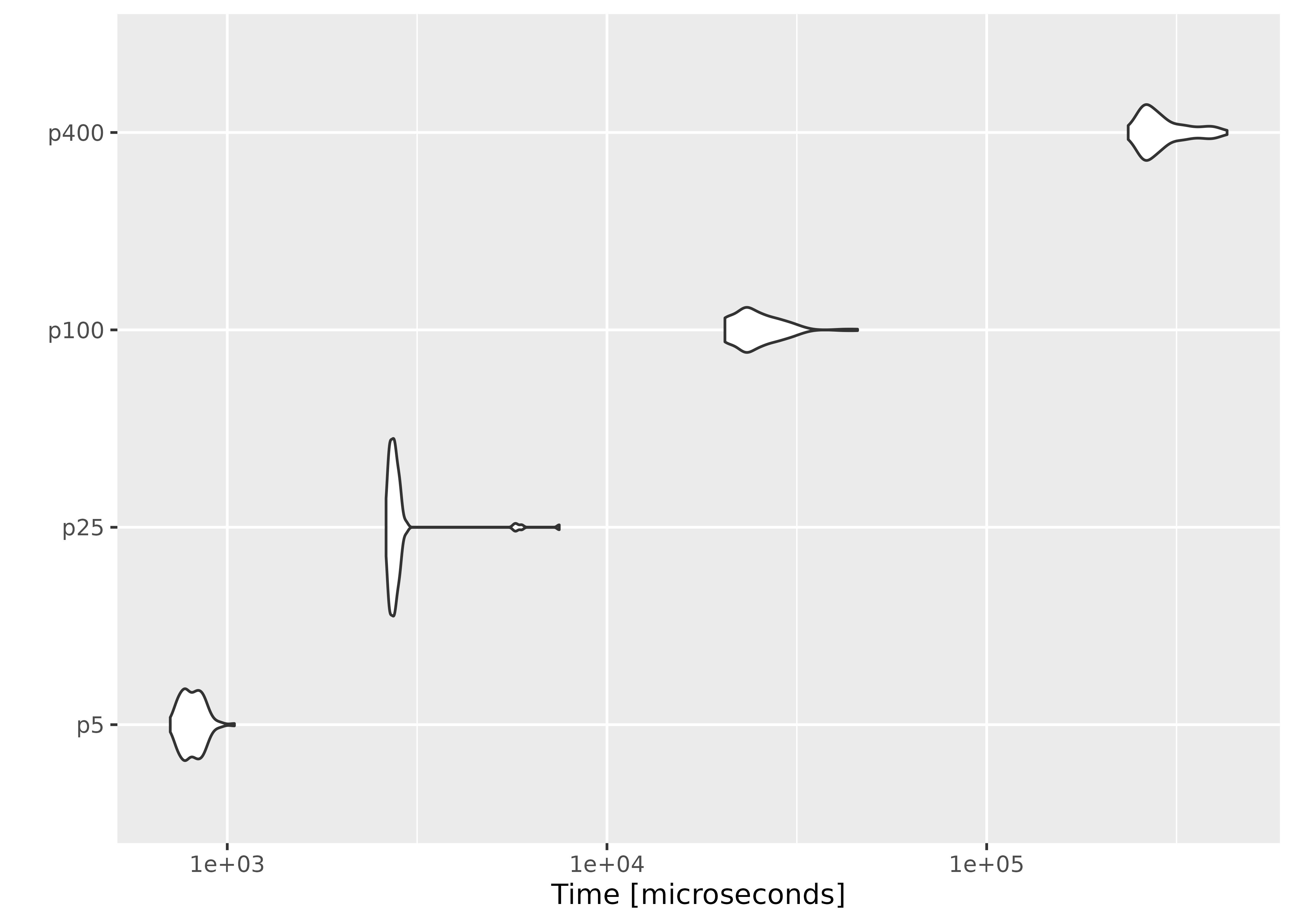
On average, evaluating empirical likelihood with a 100000×10 or 1000×400 matrix at a parameter value satisfying the convex hull constraint takes less than a second.
