europepmc facilitates access to the Europe PMC RESTful Web Service. The client furthermore supports the Europe PMC Annotations API to retrieve text-mined concepts and terms per article.
Europe PMC covers life science literature and gives access to open access full texts. Europe PMC ingests all PubMed content and extends its index with other literature and patent sources.
For more infos on Europe PMC, see:
Ferguson, C., Araújo, D., Faulk, L., Gou, Y., Hamelers, A., Huang, Z., Ide-Smith, M., Levchenko, M., Marinos, N., Nambiar, R., Nassar, M., Parkin, M., Pi, X., Rahman, F., Rogers, F., Roochun, Y., Saha, S., Selim, M., Shafique, Z., … McEntyre, J. (2020). Europe PMC in 2020. Nucleic Acids Research, 49(D1), D1507–D1514. https://doi.org/10.1093/nar/gkaa994.
Implemented API methods
This client supports the following API methods from the Articles RESTful API:
| API-Method | Description | R functions |
|---|---|---|
| search | Search Europe PMC and get detailed metadata |
epmc_search(), epmc_details(), epmc_search_by_doi()
|
| profile | Obtain a summary of hit counts for several Europe PMC databases | epmc_profile() |
| citations | Load metadata representing citing articles for a given publication | epmc_citations() |
| references | Retrieve the reference section of a publication | epmc_refs() |
| databaseLinks | Get links to biological databases such as UniProt or ENA |
epmc_db(), epmc_db_count()
|
| labslinks | Access links to Europe PMC provided by third parties |
epmc_lablinks(), epmc_lablinks_count()
|
| fullTextXML | Fetch full-texts deposited in PMC | epmc_ftxt() |
| bookXML | retrieve book XML formatted full text for the Open Access subset of the Europe PMC bookshelf | epmc_ftxt_book() |
From the Europe PMC Annotations API:
| API-Method | Description | R functions |
|---|---|---|
| annotationsByArticleIds | Get the annotations contained in the list of articles specified | epmc_annotations_by_id() |
Installation
From CRAN
install.packages("europepmc")The latest development version can be installed using the remotes package:
require(remotes)
install_github("ropensci/europepmc")Loading into R
Search Europe PMC
The search covers both metadata (e.g. abstracts or title) and full texts. To build your query, please refer to the comprehensive guidance on how to search Europe PMC: https://europepmc.org/help. Provide your query in the Europe PMC search syntax to epmc_search().
europepmc::epmc_search(query = '"2019-nCoV" OR "2019nCoV"')
#> # A tibble: 100 × 29
#> id source pmid pmcid doi title authorString journalTitle issue journalVolume
#> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 36754560 MED 36754560 PMC9… 10.1… Inno… Yerlikaya S… BMJ Open 2 13
#> 2 37400836 MED 37400836 PMC1… 10.1… Effe… Ebrahimi T,… BMC Oral He… 1 23
#> 3 37223279 MED 37223279 PMC1… 10.1… Bill… Lamsal R, R… Data Brief <NA> 48
#> 4 36727245 MED 36727245 PMC1… 10.1… Vasc… Morrissey E… JBI Evid Sy… 5 21
#> 5 37648680 MED 37648680 <NA> 10.3… [Cli… Liu ZT, Che… Zhonghua Ya… 10 59
#> 6 37211453 MED 37211453 PMC1… 10.1… Safe… Smith K, He… Vaccine 26 41
#> 7 37479685 MED 37479685 PMC1… 10.1… SuPA… Wei C, Datt… Nat Commun 1 14
#> 8 37652823 MED 37652823 <NA> 10.1… Immu… Raiser F, D… Vaccine <NA> <NA>
#> 9 37714559 MED 37714559 <NA> 10.1… New-… Kobayashi N… BMJ Case Rep 9 16
#> 10 PPR525786 PPR <NA> <NA> 10.1… The … Alihsan B, … <NA> <NA> <NA>
#> # ℹ 90 more rows
#> # ℹ 19 more variables: pubYear <chr>, journalIssn <chr>, pageInfo <chr>, pubType <chr>,
#> # isOpenAccess <chr>, inEPMC <chr>, inPMC <chr>, hasPDF <chr>, hasBook <chr>,
#> # hasSuppl <chr>, citedByCount <int>, hasReferences <chr>, hasTextMinedTerms <chr>,
#> # hasDbCrossReferences <chr>, hasLabsLinks <chr>, hasTMAccessionNumbers <chr>,
#> # firstIndexDate <chr>, firstPublicationDate <chr>, versionNumber <int>Be aware that Europe PMC expands queries with MeSH synonyms by default. You can turn this behavior off using the synonym = FALSE parameter.
By default, epmc_search() returns 100 records. To adjust the limit, simply use the limit parameter.
See vignette Introducing europepmc, an R interface to Europe PMC RESTful API for a long-form documentation about how to search Europe PMC with this client.
Creating proper review graphs with epmc_hits_trend()
There is also a nice function allowing you to easily create review graphs like described in Maëlle Salmon’s blog post:
tt_oa <- europepmc::epmc_hits_trend("Malaria", period = 1995:2019, synonym = FALSE)
tt_oa
#> # A tibble: 25 × 3
#> year all_hits query_hits
#> <int> <dbl> <dbl>
#> 1 1995 449216 1471
#> 2 1996 458644 1529
#> 3 1997 456804 1834
#> 4 1998 474693 1756
#> 5 1999 493837 1951
#> 6 2000 532142 2078
#> 7 2001 545702 2180
#> 8 2002 561497 2351
#> 9 2003 588612 2596
#> 10 2004 628176 2831
#> # ℹ 15 more rows
# we use ggplot2 for plotting the graph
library(ggplot2)
ggplot(tt_oa, aes(year, query_hits / all_hits)) +
geom_point() +
geom_line() +
xlab("Year published") +
ylab("Proportion of articles on Malaria in Europe PMC")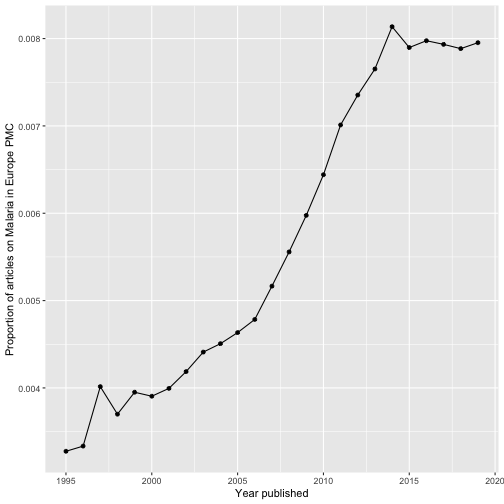
For more info, read the vignette about creating literature review graphs:
https://docs.ropensci.org/europepmc/articles/evergreenreviewgraphs.html
Re-use of europepmc
Check out the tidypmc package
https://github.com/ropensci/tidypmc
The package maintainer, Chris Stubben (@cstubben), has also created an Shiny App that allows you to search and browse Europe PMC:
Other ways to access Europe PubMed Central
Other APIs
- Data dumps: https://europepmc.org/FtpSite
- OAI service: https://europepmc.org/OaiService
- SOAP web service: https://europepmc.org/SoapWebServices
- Grants RESTful (Grist) API: https://europepmc.org/GristAPI
Other R clients
- use rOpenSci’s
oaito get metadata and full text via Europe PMC’s OAI interface: https://github.com/ropensci/oai - use rOpenSci’s
rentrezto interact with NCBI databases such as PubMed: https://github.com/ropensci/rentrez - rOpenSci’s
fulltextpackage gives access to supplementary material of open access life-science publications in Europe PMC: https://github.com/ropensci-archive/fulltext
Meta
Please note that this project is released with a Contributor Code of Conduct. By participating in this project you agree to abide by its terms.
License: GPL-3
Please use the issue tracker for bug reporting and feature requests.
